SequenceServer (BETA)
BLAST search made easy!
SequenceServer lets you rapidly set up a BLAST+ server with an intuitive user interface for use locally or over the web.
Sequence data is generated at increasing rates. However it is challenging to effectively share and query such data. Deploying complex solutions such as GMOD is often overkill and publicly available front ends for BLAST are difficult to use or install.
SequenceServer is an efficient and elegant alternative. Sequenceserver is still being developed yet is fully functional and has already been deployed for multiple databases.
SequenceServer is free for academics. Its source code is available on GitHub.
Easy for biologists
- Minimalistic, clutter-free design, so you can focus on the biology
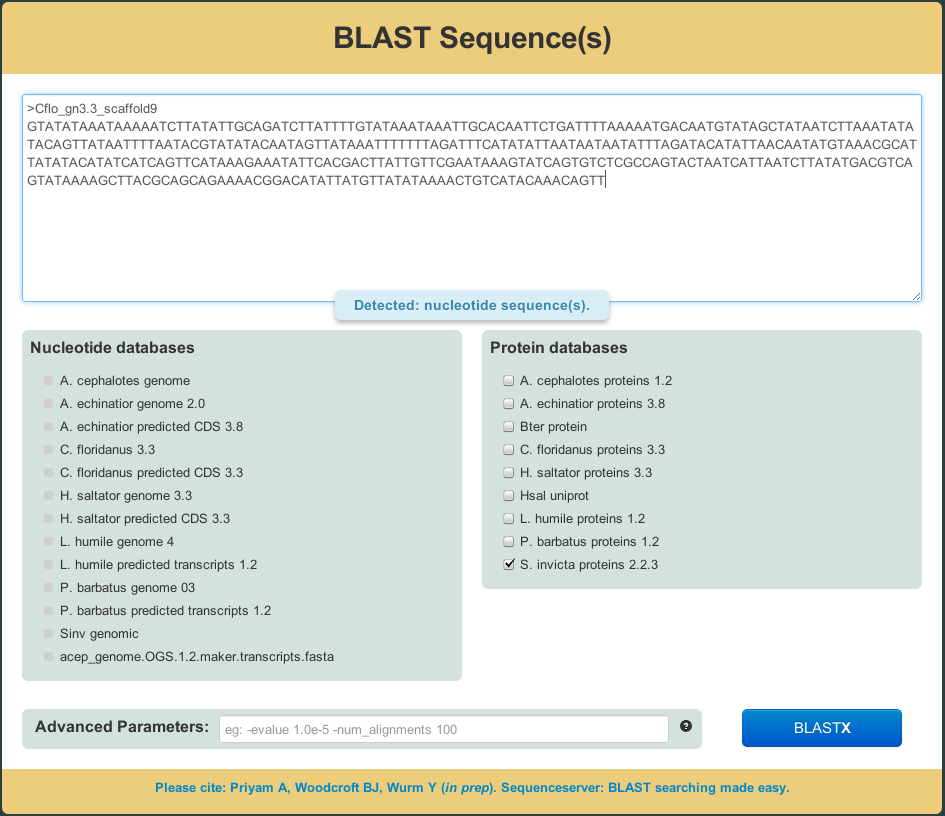
- The smart user interface automagically figures out the appropriate BLAST method to use based on your input and selected databases
- Common mistakes prevented. E.g., SequenceServer warns if you mix nucleotide and protein sequences
- Advanced users: use advanced parameters (e.g.
-task blastn-short -evalue 1.0e-20) like you would in the command line. - Simple overview of results
- Download links for hits
Rapidly deployed by admins
- very little configuration required: only a directory containing formatted or un-formatted sequence databases
- SequenceServer scans the database directory during startup. And will interactively help you create BLAST databases from FASTA files
- Comes with a built-in web server. No need to deal with Apache or Nginx to run SequenceServer.
- Easily customizable hyperlinks to search hits (e.g. to your local genome browser or custom database)
Requirements
Ruby (>= 1.8.7), RubyGems (>= 1.3.6), and NCBI BLAST+ (>= 2.2.25+). That's it!
Linux and MacOS are officially supported. While we would like to also support Windows, our resources are limited and we prefer to first concentrate on making SequenceServer great on fewer platforms.
Demo server
Sure, check Ant genome BLAST or the sites of some of our users.
Download & Setup in just a few minutes
Installation
Run the following from a command line to get the latest release of SequenceServer.
$ gem install sequenceserver
If that doesn't work, try sudo gem instead of
gem. And subsequently follow the instructions on
screen.
Configuration
SequenceServer needs to know the location of the BLAST+ binaries
(NCBI
BLAST+ download directory or
brew tap homebrew/science; brew install blast),
and the BLAST databases. This is accomplished by providing a
.sequenceserver.conf in the user's home directory.
# .sequenceserver.conf
bin: ~/ncbi-blast-2.2.25+/bin/
database: /Users/me/blast_databases/
SequenceServer automatically generates a fully commented
configuration file (.sequenceserver.conf) in the user's
home directory if it can't find one. Uncomment the relevant lines
(i.e remove the # signs), edit the values, and run
SequenceServer.
If you you already have databases, you're ready to launch SequenceServer. If not, set up databases as below.
Launch SequenceServer
Simply execute:
$ sequenceserver
SequencerServer will check your config file and start up a webserver on http://localhost:4567/. The port number can be changed in the config file.
You're done. BLAST away.
If needed: Set up custom BLAST databases
To set up BLAST databases from a directory of FASTA sequence
files, use sequenceserver's
format-databases subcommand:
$ sequenceserver format-databases
It will automatically pick up the database directory from the configuration file. Or pass the database directory from command line:
$ sequenceserver format-databases directory_with_fasta_files
The script will assist you by:
- finding FASTA files in
directory_with_fasta_files(and in subdirectories too) - detecting the sequence type of each file
- asking you to name each database
- subsequently running the appropriate
makeblastdbcommands.
Alternatively, manually set up BLAST databases from single FASTA sequence files. In the directory containing the FASTA file, run:
$ makeblastdb -dbtype <db type> -title <db title> -in <db> -parse_seqids
Where,
<db type>is eitherprot, ornucldepending on the type of sequence<db title>is what users will see<db>is the path to the FASTA file-parse_seqidsis required to generate links for downloading search hits (yes, it's a bit slow on large files).- Additional options at
makeblastdb -help.
Optional: SequenceServer on Apache or Nginx with Phusion Passenger
For most people, simply running the SequenceServer as above is more than enough. But if you need to deploy SequenceServer BLAST as part of a standard website, and have it show up as a normal webdirectory, SequenceServer can be deployed on Apache or Nginx with Phusion Passenger™ (a.k.a mod_rails or mod_rack).
Set up Phusion Passenger™
Install passenger gem:
gem install passenger
Install passenger module for Apache/Nginx module
# for apache2
passenger-install-apache2-module
# for nginx
passenger-install-nginx-module
Follow instructions on screen. The installer will let you know of
any missing software that is required to setup passenger, and also
how to install them. Towards the end of installation, after the
installer has finished building modules corresponding to your
webserver, it will ask you to edit your webserver configuration files
and some lines, so that your webserver knows how to load
passenger.
For Apache, you can do something like this:
- Create a new file called
passenger.loadin/etc/apache2/mods-available, and add the lines there. - Run
a2enmod passenger. This will ask you to restart apache2 with instructions on how to do the same. On my machine, I had to run/etc/init.d/apache2 restart - Running
touch tmp/restart.txtmay required when changing configuration.
Deploying SequenceServer
Apache Root URI (eg:antgenomes.org or localhost).
<VirtualHost *:80>
ServerName localhost
DocumentRoot /home/yeban/src/sequenceserver/public
<Directory /home/yeban/src/sequenceserver/public>
Allow from all
Options -MultiViews
</Directory>
</VirtualHost>
Apache Sub URI (e.g.: antgenomes.org/blast or localhost/sequenceserver).
<VirtualHost *:80>
ServerName localhost
DocumentRoot /var/www/
<Directory /var/www>
Allow from all
</Directory>
RackBaseURI /sequenceserver
<Directory /var/www/sequenceserver>
Options -MultiViews
</Directory>
</VirtualHost>
Support Mailing List
Have an issue in deploying SequenceServer? Something is not working as expected? Have a tip? A feature request? Or just want to encourage further development? Post it to SequenceServer Google Group and we will work something out.
If you are completely overwhelmed and need more support that the mailing list can provide, we are available for consulting which can range from custom support to server deployment and administration to implementing specific features.
License
For academics and non-for-profit organizations
SequenceServer is free & you can do most things you want with it. Our License.txt file details some constraints regarding use and re-distribution (e.g. that instances you install retain links to this website). Please remember to cite the software.For Commercial & for-profit organizations
If you're a for-profit you can try it with no constraints for one month. Subsequently you must purchase a license (it's cheap, quick & easy!).Custom licensing is also possible. Get in touch.
FAQ Frequently Asked Questions
- How do I cite SequenceServer?
- A manuscript describing SequenceServer is in prep. In the mean time, please cite the following reference: "Priyam A., Woodcroft B.J., Wurm Y in prep. SequenceServer: BLAST searching made easy". This has already been done several times.
- What are the alternatives to SequenceServer?
- There are several. But they are suboptimal because they look like they're from the last millenium. And installing them is challenging.
-
- NCBI's wwwblast hasn't been maintained in years and is difficult to set up.
- ViroBLAST requires Apache & MySQL.
- GMOD's BLAST Graphic Viewer hasn't been maintained in years.
- BLASTstation is only for local BLAST.
- Is there a demo server?
- SequenceServer provides BLAST for ant genomes. Some of our users also provide publicly accessible servers.
- What is a BLAST database? Can I make a custom BLAST database?
- The BLAST program doesn't understand FASTA files. But BLAST
includes the
makeblastdbtool that reformats FASTA into the optimized BLAST-friendly format. Usemakeblastdborsequenceserver format-databasesto create a custom BLAST database. - Can I use a preformatted BLAST database? Can I use alias?
- Preformatted BLAST databases can also be used with SequenceServer. Most preformatted database ar split between multiple files (like .00, .01...). Thanks to Mark Anthony Gibbins' contributions, SequenceServer can correctly understand these.
- If you use a custom
.nalor.palBLAST database alias file, and copy this alias file as well as the linked databases into SequenceServer'sdbdirectory, SequenceServer will display both the alias and the linked databases. If you want to display only the alias, remove the linked databases fromdb, then edit theDBLISTline in the.palor.nalfile so that it contains the complete path of each of the database files (e.g:DBLIST species1_custom_blast_db species2_custom_blast_databasemay become:DBLIST /Users/blastuser/mydbs/species1_custom_blast_db /Users/blastuser/mydbs/species2_custom_blast_database) - How does BLAST identify similar regions? Why don't the ends match? What is the bitscore? What is the e-value that BLAST returns? What do these numbers mean?
- Many detailed explanations exist egarding such questions, including at NCBI and in the original BLAST articles. Here's our executive summary.
- BLAST is a heuristic, ie. it is fast and approximate instead of being slow and perfect. It starts by looking for a minimal 100% match (e.g. 11 consecutive nucleotides with 100% identity between your query and the database sequence). It is done if it finds none (thus if exactly every 10th base is different, BLAST finds no results). If it does find a match, it extends that in both directions: identical (or similar) bases add points; differences are negative points. If too many points are lost, it stops aligning. It might not stop at the exact best place, thus ends might not match. The "bitscore" is the total number of points for the aligning region. The bigger it is, the stronger the alignment. But the bitscore doesn't take into account sequence length nor database size. The E-value does. It is better to look at E-values than bitscores. The E-value represents the number of times the observed alignment would be expected to occur by chance (it is not a p-value!); depends on the bitscore, the length of the query sequence, and the cumulative length of all sequences in the database. Its easier to talk about "strong" E-values (e.g. 1e-100 = 10-100 = almost zero; impossible to obtain by chance) vs "weak" E-evalues (e.g 0.1; for similarity that may be due to chance) that small vs large (which is always a bit confusing).
- What is the difference between BLAST and BLAST+? What about WU-BLAST or AB-BLAST?
- The initial BLAST has been rewritten several times - most recently by NCBI as BLAST+. NCBI now use and recommend using BLAST+. The BLAST+ publication explains why BLAST+ is easier to use & faster than the old "legacy" BLAST. WU-BLAST is now commerical and called AB-BLAST. There is probably no good reason to use either. Note that the output formats change slightly from one BLAST implementation to the next.
- NCBI's BLAST+ is most actively developed, just use it. SequenceServer only supports BLAST+.
- I swear I installed BLAST but SequenceServer can't find it!
- Make sure you downloaded NCBI's Blast+ (Blast Plus!) and not legacy BLAST. The old legacy
BLAST had programs such as
blastallandformatdb. The programs in BLAST+ have names likemakeblastdb,blastn,tblastx... - Also, check that your SequenceServer configuration file points at the
bindirectory, e.g.:/Users/stevejobs/software/ncbi-blast-2.2.25+/bin/ - Why doesn't sequence retrieval work?
- Probably because you forgot the
-parse_seqidsargument when runningmakeblastdb(or they were formatted using legacyformatdb). Blast+ 2.2.24 and later indicate that when-parse_seqidswas not used by leaving a space between>and hit identifiers (eg:> db_sequence001instead of>db_sequence001). - How can I add custom links to hits?
- Easy. Edit the
customisation.rbfile and read the examples it provides. To add a single link, uncomment and edit theconstruct_custom_sequence_hyperlinkmethod; to add multiple links edit theconstruct_custom_sequence_hyperlinking_linemethod. - How do I tell SequenceServer to read a different configuration file?
- By using
--configcommand line switch. For example:$ sequenceserver --config sequenceserver_ants.conf
You will need to set it inconfig.rufor Rack based deployment:SequenceServer::App.config_file = 'sequenceserver_ants.conf'
The above line should be added beforeSequenceServer::App.init, so theconfig.rulooks something like this:require 'rubygems' require 'bundler/setup' require 'sequenceserver' SequenceServer::App.config_file = '/absolute/path/to/configuration_file' SequenceServer::App.init run SequenceServer::App
- How do I change the server port?
- SequenceServer will by default be accessible on port 4567
at http://localhost:4567/.Change
the port by editing
~/.sequenceserver.conf. - Can my colleagues access the server I set up on my computer?
- Yes, if they can access your machine. This usually requires being in the same subnetwork, or asking IT services to open your machine to the outside world. You'll have to tell your colleagues your IP address (find it in the Network or Sharing section of System Preferences) and the port. The complete URL they will have to use might be http://192.168.134.13:4567/ or http://myblastserver.local:4567/
- Do I need to install Apache?
- SequenceServer automatically uses ruby's internal web servers. No
need for Apache. By default it uses Webrick.
Installling Thin
(
gem install thin), or Mongrel (gem install mongrel) can improve performance. SequenceServer will automatically use these if installed. - Can I integrate SequenceServer into an Apache or Nginx server?
- Yes. We provide instructions above.
- Should I activate hyper-threading to give BLAST more virtual cores?
- Yes.
- Which technologies does SequenceServer use?
- SequenceServer uses NCBI BLAST+, Sinatra, and jQuery.
- Can I have the source code? Can I contribute?
- SequenceServer's source code is on GitHub. Downloading, forking, pulling, filing issues and contributing is most welcome!
- You say SequenceServer is easy to set up. But I'm not a computer person and I don't understand a thing. This is too hard!
- We are considering offering hosted (password-protected) servers for a fee (ie you upload the data; we set things up as you need them). Please get in touch by email if you are interested or need other consulting advice.
- Can I get a support contract which guarantees rapid response?
- Yes. Get in touch by email.
- Can SequenceServer do XXX? Why doesn't YYYY work? It's ugly, it crashes!?!?
- First, run
gem updateorsudo gem update. - Feel free to add comments, vote, report bugs, request features and contribute code. All contributions, patches and pulls are welcome. Our issues tracker logs development activities. If you're not a coder, please consider supporting development by a donation. Donations can dramatically increase the speed at which new features are added!
Contributors
- Anurag Priyam
- is a software developer primarily interested in applied computing and web infrastructure development. He is currently pursuing his undergraduate studies at the Indian Institute of Technology, Kharagpur.
- Ben J Woodcroft
- is researching microbial responses to permafrost thaw (climate change induced), and the bioinformatics of metagenomics. He is a post-doctoral fellow at the Australia Centre for Ecogenomics.
- Yannick Wurm
- is an evolutionary biologist using genomics tools to study the evolution of ant societies (eg: antgenomes.org, fire ant genome, competing fire ant queens). He is a lecturer at Queen Mary University London.
- Maybe you?
- The SequenceServer team will happily accept contributions. Please get in touch to figure out where you could fit in. Cheers!
Users include:

As well as:
- AvianGenomes.org
- iPlantCollaborative Saccharum Genome Database
- BioCloud.ca
- The Streptocarpus rexii database at Bioinformatics Evolution and COmparative Genomics lab, Uni Milan
- Dr. Wolfang Rumpf @ U Maryland
- Center for Ecological Research, Kyoto University
- European clam database @ CCMAR at University of Algarve
- Spotted Drosophila Flybase @ Oregon State
- Setariabase for Setaria viridis at Donald Danforth Plant Science Center
- at Open Ash Dieback GeeFu
- Amborella Genome at University of Georgia
- Jekely Lab at Max Planck Institute for Developmental Biology, Tuebigen
- Universitat Greifswald (private)
- Department of Plant Biology and Forest Genetics, Swedish University of Agricultural Sciences (private)
- Norwich BioScience Institutes (private)
- Australia's Commonwealth Scientific and Industrial Research Organisation (private)
- Max Planck Institute of Molecular Cell Biology and Genetics (private)
- School of Biological Sciences, University of Queensland (private)
and many additional private installations we don't know about...
Citations include:
- Mondav R et al (2014) Discovery of a novel methanogen prevalent in thawing permafrost Nature Communications 5, 3212.
- Rowe ML et al (2014) Neuropeptides and polypeptide hormones in echinoderms: New insights from analysis of the transcriptome of the sea cucumber Apostichopus japonicus General and Comparative Endocrinology 194, 43-55.
- Chiara M et al (2013) De Novo Assembly of the Transcriptome of the Non-Model Plant Streptocarpus rexii Employing a Novel Heuristic to Recover Locus-Specific Transcript Clusters PLoS ONE.
- Semmens DC et al (2013) Discovery of a novel neurophysin-associated neuropeptide that triggers cardiac stomach contraction and retraction in starfish Journal of Experimental Biology 216, 4047-4053.
- Chiu JC et al (2013) Genome of Drosophila suzukii, the Spotted Wing Drosophila G3 g3.113.008185
- Elphick MR (2012) The Protein Precursors of Peptides That Affect the Mechanics of Connective Tissue and/or Muscle in the Echinoderm Apostichopus japonicus. PLoS ONE.
- Shreve J et al (2013) A genome-wide survey of small interfering RNA and microRNA pathway genes in a galling insect. Journal of Insect Physiology.
- Berlamino et al (2013) SymGRASS: a database of sugarcane orthologous genes involved in arbuscular mycorrhiza and root nodule symbiosis. BMC Bioinformatics.
- Elphick MR et al (2013) The Evolution and Diversity of SALMFamide Neuropeptides. PLoS ONE.
Parts of this software are © 2011 University of Lausanne.
Please cite: "Priyam A., Woodcroft B.J., Wurm Y in prep. SequenceServer: BLAST
searching made easy http://sequenceserver.com."